Derda Lab bridges DNA-Encoded Chemistry and Machine Learning to solve fundamental problems in chemical reactivity, chemical biology, molecular recognition and drug discovery”
We develop DNA-encoded ligand discovery and fragment-based discovery with the goal of developing drug candidates, Targeted Radionuclide Therapies, and targeting vectors for the delivery of next-generation of genetic medicines such as RNA-therapeutics and CRISPR/Cas9 therapeutics. We employ DNA-encoded billion-scale molecular libraries developed by our group to solve the “impossible problems in molecular recognition”. One of such problems is identification of ligands for “undruggable targets”: these are molecular receptors for which there are few or no therapeutics available. Proteins that recognize carbohydrates are one of the examples of such targets; we disclosed examples of discoveries of molecules that block protein-carbohydrate interactions. Our DNA-encoded discovery pipeline has been commercialized by 48Hour Discovery Inc.
The fundamental goal of this research is to employ information provided by DNA-encoded libraries to train Machine Learning / AI models to predict molecular interactions. Automation of molecular discoveries by generative AI approaches has been reported but all modern AI approaches require some data already to exist in order for these algorithms to be successful. This fundamental “cold start problem” can be solved with the help of DNA-encoded libraries, which provide the critical data to “jump start” the AI-driven discovery engine. A successful combination of DNA-encoded libraries and AI technologies will make it possible to develop automated molecular discoveries for first-in-class targets (i.e., targets for which there is presently no information available).
1. Derda and Ng “Genetically-Encoded Fragment-Based Discovery (GE-FBD)”, Curr. Op. in Chemical Biology, 2019, 50, 128.
2. Ng, et al, “Genetically-encoded Fragment-based Discovery of Glycopeptide Ligands for Carbohydrate-binding Proteins“, JACS 2015, 137, 5248.
3. Tjhung et. al. “Silent Encoding of Chemical Post-Translational Modifications in Phage-Displayed Libraries“, JACS 2016, 138, 32
4. Ekanayake, et al. “Genetically Encoded Fragment-Based Discovery (GE-FBD) from Phage-Displayed Macrocyclic Libraries with Genetically-Encoded Unnatural Pharmacophores” JACS 2021 143, 14, 5497.
5. Yan et al. “Late-Stage Reshaping of Phage-Displayed Libraries to Macrocyclic and Bicyclic Landscapes using a Multipurpose Linchpin” JACS 2025, 147, 1, 789
6. Alteen, et. al. “Phage display uncovers a sequence motif that drives polypeptide binding to a conserved regulatory exosite of O-GlcNAc transferase” PNAS, 2023, 120, e2303690120.
DNA-encoded libraries of chemicals make it possible to run millions of reaction in parallel and analyze their outcomes by Next-Generation Sequencing (NGS). Linking chemical reactivity to DNA barcodes makes it possible to amplify the outcomes from rare reaction events using biological amplification (e.g., PCR). We demonstrated that DNA-libraries of peptides displayed on phage can be successfully employed in a diverse range of reactions. This platform is compatible with substrate profiling and reaction discovery with in million-scale library and machine learning of new chemical transformation.
Our long term goal is to use DNA-encoding and DNA-amplification change the paradigm in discovery and optimization of chemical reactions, chemical catalysis and enzyme-driven chemical transformation.
1. Yan et al “Learning Structure Activity Relationship (SAR) of the Wittig Reaction from Genetically-Encoded Substrates“, Chem. Sci., 2021,12, 14301-14308.
2. Kalhor-Monfared et al. “Rapid Biocompatible Macrocyclization of Peptides with Decafluoro-diphenylsulfone” Chem. Sci., 2016, 7, 3785-3790.
2. Kitov et al “Rapid, Hydrolytically Stable Modification of Aldehyde-terminated Proteins and Phage Libraries”, J. Am. Chem. Soc., 2014, 136, 8149–8152.
4. Triana and Derda, “Tandem Wittig / Diels-Alder Diversification of Genetically Encoded Peptide Libraries“, Org. Biomol. Chem., 2017, 15, 7869-7877
5. Ng, Jafari and Derda “Bacteriophages and Viruses as a Support for Organic Synthesis and Combinatorial Chemistry“, ACS Chem. Biol. 7, 123.
.
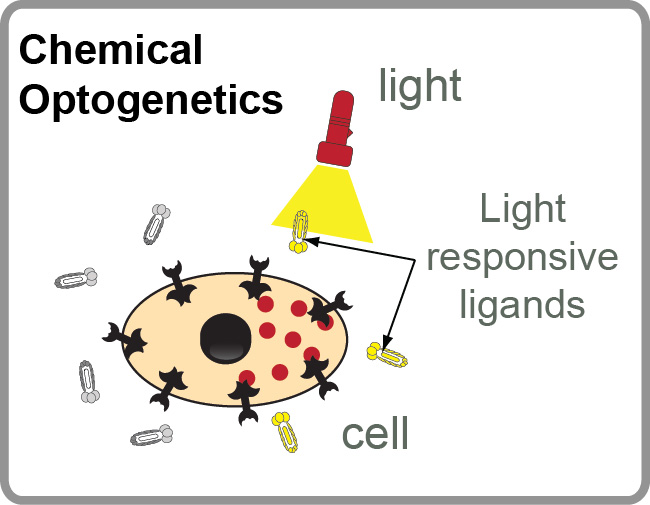
The filed of “photopharmacology” aims to develop therapeutics compounds that can be activated by light in desired tissue, cell or sub-cellular location at a specific time. “Chemical optogenetics” is a field on the interface of “chemical genetics” (interrogation of cell function using small-molecule inhibitors) and “optogenetics” (investigation of cell functions using proteins that can be reversibly activated by light). The main challenge in both fields is discovery of light-responsive molecules that can be reversibly turned ON and OFF by light. To address this challenge, we are developing methods for DNA-encoded discovery of molecules that can be reversibly activated or deactivated with light.
1. MR Jafari, Lu Deng, S Ng, W Matochko, K Tjhung, A Zeberof, A Elias, John S. Klassen, R Derda* “Discovery of light-responsive ligands through screening of light-responsive genetically-encoded library“, ACS Chem. Biol., 2014, 9, 443–450
2. M. R. Jafari, J. Lakusta, R. J. Lundgren, and R Derda* “Allene Functionalized Azobenzene Linker Enables Rapid and Light-Responsive Peptide Macrocyclization” Bioconj. Chem., 2016, 27, 509–514
3. Mohammad R. Jafari, Hongtao Yu, Jessica M. Wickware, Yu-Shan Lin and Ratmir Derda*, “Light Responsive Bicyclic Peptides” Org. Biomol. Chem., 2018,16, 7588-7594.
Forward chemical genetics identifies molecules that control any cellular phenotypes. Billion-scale DNA-encoded libraries are uniquely positioned for such discovery. Combining stem cells with such DNA-encoded libraries can discover molecules that control differentiation of such cells. This powerful approach requires no knowledge about the structure of molecular receptors nor signalling pathways controlled by such receptors. We combine DNA-encoded library, molecular array technologies and other approaches to develop chemical genetic approaches capable of unsupervised discovery of molecules and materials that control differentiation of cells.
1. Derda et al. “High-throughput Discovery of Synthetic Surfaces that Support Proliferation of Pluripotent Cells” JACS 2010.
2. Deiss, et al., “Flow-Through Synthesis on Teflon-Patterned Paper To Produce Peptide Arrays for Cell-Based Assays”, Angewandte Chemie, 2014, 53, 6374.
3. Lin, et al., “Peptide Microarray Patterning for Controlling and Monitoring Cell Growth” Acta Biomater., 2016, 34, 53.
4. Matochko et al. “Reproducible Discovery of Cell-Binding Peptides ‘Lost’ in Bulk Amplification via Emulsion Amplification in Phage Display Panning“. BioRvix 2021, DOI 2021.10.31.466683v1
5. Derda et al. “Paper-Supported Three-Dimensional Cell Culture for Tissue-Based Bioassays” PNAS 2009, 106, 18457.
Rapid search in vast chemical space can help solve many fundamental problems in molecular recognition and chemical reactivity. However, like any large-scale information source (e.g., the Internet), search in chemical space requires an efficient “search engine” and a fundamental understanding of “discovery trajectories”. Our group aims to understand the fundamental forces that govern molecular discovery trajectories and the reproducibility of molecular discoveries. Like the Internet, the vast chemical space of genetically encoded libraries contains areas that are censored and cannot be easily accessed. Biased and unbiased molecular discovery can be compared to searching the Internet using biased and bias-free search engines. We employ deep-sequencing technology to characterize these biases, such as “parasitic sequences” in molecular discovery. Our knowledge clouds store thousands of such discovery trajectories and employ them to train Machine Learning algorithms for effective search. We demonstrated that micro-droplets remove unwanted biases in discovery campaigns by ceasing unwanted competition between members of DNA-encoded libraries. Our group introduced fundamental concepts into molecular discovery such as “discovery baseline” and “discovery operators”, with the long-term goal of automating molecular discoveries.
1. Yan, et al, “Universal Baseline for in vitro Selection of Genetically Encoded Libraries” BioRxiv 2026
2. Derda et al. “Diversity of Phage-Displayed Libraries of Peptides During Panning and Amplification” Molecules 2011, 16, 1776-1803
3. (a) Derda et al. “Uniform Amplification of Phage with Different Growth Characteristics in Monodisperse Droplet-Based Compartments” Angew. Chem. Intl. Ed. 2010 49, 5301; (b) Matochko et al. “Uniform amplification of phage display libraries in monodisperse emulsions“, Methods, 2012, 58, 18.
4. Matochko et al. “Prospective identification of parasite sequences in phage display screens” Nuc. Acid. Res., 2014, 42, 17845.
5. Matochko et al. “Reproducible Discovery of Cell-Binding Peptides ‘Lost’ in Bulk Amplification via Emulsion Amplification in Phage Display Panning“. BioRvix 2021
6. Matochko et al. “Error Analysis of Deep Sequencing of Phage Libraries: Peptides Censored in Sequencing” Comput Math Methods Med. 2013
The Central Dogma of Biology, DNA → RNA → Protein, facilitates the study of biology using a unified toolbox of DNA sequencing. This ability has revolutionized and transformed all areas of biomedical and life science. Carbohydrates are one of the most diverse and ubiquitous components in all kingdoms of life, but investigating the biological roles of carbohydrates cannot rely on DNA sequencing directly. To solve this problem, we developed a technology that introduces a one-to-one correspondence between DNA sequence and carbohydrate structure [1]. Genetically encoded multivalent Liquid Glycan Array (LiGA) and Liquid Lectin Array (LiLA) enable investigation of glycobiology using DNA sequencing, single-cell RNA sequencing, and spatial transcriptomics. A major advantage of “liquid arrays” is the ability to measure protein:carbohydrate interactions in specific cells in live organisms in vivo. The long-term fundamental goal is to combine array technologies with machine learning to solve the fundamental long-standing problem of the “glycan code”: given the atomic structure of a glycan and the amino acid sequence of a protein, predict how strongly this glycan and this protein interact with one another.
1. Sojitra et al. “Genetically encoded multivalent liquid glycan array displayed on M13 bacteriophage“, Nature Chem. Biol (2021).
2. Lin et al. “Chemoenzymatic synthesis of genetically-encoded multivalent liquid N-glycan arrays” Nature Comm. 14, 5237 (2023)
3. Sojitra et al. “Measuring carbohydrate recognition profile of lectins on live cells using liquid glycan array (LiGA)” Nature Prot. (2024)
4. Lima, et al. “The liquid lectin array detects compositional glycocalyx differences using multivalent DNA-encoded lectins on phage“, Cell Chem. Biol. (2024)
5. Carpenter et al. “GlyNet: A Multi-Task Neural Network for Predicting Protein-Glycan Interactions“, Chem. Science, (2022).
6. Carpenter et al. “Atom-level machine learning of protein-glycan interactions and cross-chiral recognition in glycobiology” Science Adv. 2025.